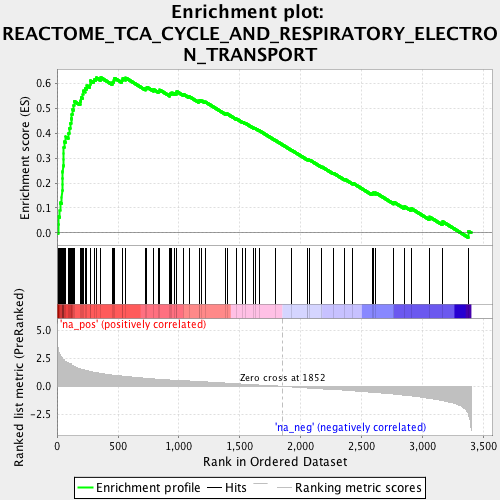
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | chrom |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | REACTOME_TCA_CYCLE_AND_RESPIRATORY_ELECTRON_TRANSPORT |
| Enrichment Score (ES) | 0.6218272 |
| Normalized Enrichment Score (NES) | 2.4524662 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | NDUFA4 | NDUFA4 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 4, 9kDa | 9 | 3.447 | 0.0330 | Yes |
| 2 | NDUFS4 | NDUFS4 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 4, 18kDa (NADH-coenzyme Q reductase) | 12 | 3.194 | 0.0655 | Yes |
| 3 | NDUFA6 | NDUFA6 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 6, 14kDa | 23 | 2.843 | 0.0919 | Yes |
| 4 | NDUFS6 | NDUFS6 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 6, 13kDa (NADH-coenzyme Q reductase) | 25 | 2.804 | 0.1206 | Yes |
| 5 | NDUFV2 | NDUFV2 Entrez, Source | NADH dehydrogenase (ubiquinone) flavoprotein 2, 24kDa | 39 | 2.536 | 0.1430 | Yes |
| 6 | NDUFA7 | NDUFA7 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 7, 14.5kDa | 40 | 2.525 | 0.1691 | Yes |
| 7 | COX6B1 | COX6B1 Entrez, Source | cytochrome c oxidase subunit Vib polypeptide 1 (ubiquitous) | 43 | 2.496 | 0.1943 | Yes |
| 8 | NDUFA2 | NDUFA2 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 2, 8kDa | 44 | 2.444 | 0.2197 | Yes |
| 9 | NDUFB7 | NDUFB7 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 7, 18kDa | 46 | 2.423 | 0.2444 | Yes |
| 10 | NDUFA12 | NDUFA12 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 12 | 49 | 2.411 | 0.2688 | Yes |
| 11 | NDUFS1 | NDUFS1 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 1, 75kDa (NADH-coenzyme Q reductase) | 53 | 2.397 | 0.2927 | Yes |
| 12 | NDUFA10 | NDUFA10 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 10, 42kDa | 54 | 2.384 | 0.3174 | Yes |
| 13 | COX5B | COX5B Entrez, Source | cytochrome c oxidase subunit Vb | 55 | 2.381 | 0.3421 | Yes |
| 14 | NDUFB9 | NDUFB9 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 9, 22kDa | 60 | 2.305 | 0.3647 | Yes |
| 15 | NDUFB1 | NDUFB1 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 1, 7kDa | 71 | 2.219 | 0.3847 | Yes |
| 16 | NDUFB8 | NDUFB8 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 8, 19kDa | 93 | 2.088 | 0.4000 | Yes |
| 17 | NDUFB6 | NDUFB6 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 6, 17kDa | 100 | 2.065 | 0.4196 | Yes |
| 18 | NDUFV1 | NDUFV1 Entrez, Source | NADH dehydrogenase (ubiquinone) flavoprotein 1, 51kDa | 108 | 2.033 | 0.4385 | Yes |
| 19 | BSG | BSG Entrez, Source | basigin (Ok blood group) | 115 | 1.992 | 0.4573 | Yes |
| 20 | NDUFS5 | NDUFS5 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 5, 15kDa (NADH-coenzyme Q reductase) | 121 | 1.913 | 0.4756 | Yes |
| 21 | NDUFB3 | NDUFB3 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 3, 12kDa | 127 | 1.866 | 0.4934 | Yes |
| 22 | NDUFA3 | NDUFA3 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 3, 9kDa | 136 | 1.783 | 0.5095 | Yes |
| 23 | UQCRC2 | UQCRC2 Entrez, Source | ubiquinol-cytochrome c reductase core protein II | 140 | 1.775 | 0.5270 | Yes |
| 24 | NDUFV3 | NDUFV3 Entrez, Source | NADH dehydrogenase (ubiquinone) flavoprotein 3, 10kDa | 190 | 1.570 | 0.5285 | Yes |
| 25 | NDUFA9 | NDUFA9 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 9, 39kDa | 197 | 1.525 | 0.5424 | Yes |
| 26 | NDUFB4 | NDUFB4 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 4, 15kDa | 212 | 1.489 | 0.5536 | Yes |
| 27 | NDUFS3 | NDUFS3 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 3, 30kDa (NADH-coenzyme Q reductase) | 214 | 1.482 | 0.5687 | Yes |
| 28 | COX6C | COX6C Entrez, Source | cytochrome c oxidase subunit VIc | 230 | 1.430 | 0.5790 | Yes |
| 29 | DLD | DLD Entrez, Source | dihydrolipoamide dehydrogenase | 245 | 1.397 | 0.5892 | Yes |
| 30 | COX7A2L | COX7A2L Entrez, Source | cytochrome c oxidase subunit VIIa polypeptide 2 like | 270 | 1.348 | 0.5960 | Yes |
| 31 | NDUFC2 | NDUFC2 Entrez, Source | NADH dehydrogenase (ubiquinone) 1, subcomplex unknown, 2, 14.5kDa | 273 | 1.345 | 0.6093 | Yes |
| 32 | LDHA | LDHA Entrez, Source | lactate dehydrogenase A | 303 | 1.258 | 0.6136 | Yes |
| 33 | UQCRC1 | UQCRC1 Entrez, Source | ubiquinol-cytochrome c reductase core protein I | 320 | 1.221 | 0.6214 | Yes |
| 34 | NDUFB5 | NDUFB5 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 5, 16kDa | 359 | 1.148 | 0.6218 | Yes |
| 35 | NDUFB10 | NDUFB10 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 10, 22kDa | 454 | 0.996 | 0.6038 | No |
| 36 | NDUFS8 | NDUFS8 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 8, 23kDa (NADH-coenzyme Q reductase) | 467 | 0.984 | 0.6104 | No |
| 37 | PDHX | PDHX Entrez, Source | pyruvate dehydrogenase complex, component X | 470 | 0.981 | 0.6200 | No |
| 38 | NDUFA11 | NDUFA11 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 11, 14.7kDa | 533 | 0.910 | 0.6107 | No |
| 39 | UQCRFS1 | UQCRFS1 Entrez, Source | ubiquinol-cytochrome c reductase, Rieske iron-sulfur polypeptide 1 | 540 | 0.907 | 0.6183 | No |
| 40 | PDK3 | PDK3 Entrez, Source | pyruvate dehydrogenase kinase, isozyme 3 | 564 | 0.877 | 0.6204 | No |
| 41 | UQCRB | UQCRB Entrez, Source | ubiquinol-cytochrome c reductase binding protein | 726 | 0.716 | 0.5793 | No |
| 42 | NDUFS7 | NDUFS7 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 7, 20kDa (NADH-coenzyme Q reductase) | 738 | 0.704 | 0.5833 | No |
| 43 | UQCRQ | UQCRQ Entrez, Source | ubiquinol-cytochrome c reductase, complex III subunit VII, 9.5kDa | 793 | 0.661 | 0.5739 | No |
| 44 | FH | FH Entrez, Source | fumarate hydratase | 835 | 0.629 | 0.5681 | No |
| 45 | NDUFS2 | NDUFS2 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 2, 49kDa (NADH-coenzyme Q reductase) | 843 | 0.620 | 0.5724 | No |
| 46 | ATP5C1 | ATP5C1 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, gamma polypeptide 1 | 927 | 0.562 | 0.5532 | No |
| 47 | NDUFA13 | NDUFA13 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 13 | 931 | 0.561 | 0.5581 | No |
| 48 | CYC1 | CYC1 Entrez, Source | cytochrome c-1 | 943 | 0.556 | 0.5606 | No |
| 49 | OGDH | OGDH Entrez, Source | oxoglutarate (alpha-ketoglutarate) dehydrogenase (lipoamide) | 967 | 0.540 | 0.5592 | No |
| 50 | NDUFA8 | NDUFA8 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 8, 19kDa | 981 | 0.534 | 0.5608 | No |
| 51 | SDHB | SDHB Entrez, Source | succinate dehydrogenase complex, subunit B, iron sulfur (Ip) | 983 | 0.532 | 0.5660 | No |
| 52 | COX4I1 | COX4I1 Entrez, Source | cytochrome c oxidase subunit IV isoform 1 | 1038 | 0.504 | 0.5550 | No |
| 53 | NDUFA5 | NDUFA5 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 5, 13kDa | 1085 | 0.471 | 0.5460 | No |
| 54 | ATP5L | ATP5L Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit G | 1166 | 0.420 | 0.5263 | No |
| 55 | ETFDH | ETFDH Entrez, Source | electron-transferring-flavoprotein dehydrogenase | 1167 | 0.419 | 0.5306 | No |
| 56 | ATP5F1 | ATP5F1 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit B1 | 1188 | 0.409 | 0.5288 | No |
| 57 | ATP5J2 | ATP5J2 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit F2 | 1217 | 0.399 | 0.5245 | No |
| 58 | ATP5O | ATP5O Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, O subunit (oligomycin sensitivity conferring protein) | 1387 | 0.291 | 0.4766 | No |
| 59 | ATP5B | ATP5B Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, beta polypeptide | 1396 | 0.283 | 0.4771 | No |
| 60 | ATP5H | ATP5H Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit d | 1473 | 0.233 | 0.4567 | No |
| 61 | ATP5A1 | ATP5A1 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, alpha subunit 1, cardiac muscle | 1526 | 0.193 | 0.4430 | No |
| 62 | ATP5J | ATP5J Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit F6 | 1548 | 0.184 | 0.4386 | No |
| 63 | PDHB | PDHB Entrez, Source | pyruvate dehydrogenase (lipoamide) beta | 1614 | 0.143 | 0.4205 | No |
| 64 | IDH2 | IDH2 Entrez, Source | isocitrate dehydrogenase 2 (NADP+), mitochondrial | 1627 | 0.131 | 0.4182 | No |
| 65 | COX5A | COX5A Entrez, Source | cytochrome c oxidase subunit Va | 1661 | 0.114 | 0.4095 | No |
| 66 | ATP5D | ATP5D Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, delta subunit | 1790 | 0.037 | 0.3713 | No |
| 67 | ETFB | ETFB Entrez, Source | electron-transfer-flavoprotein, beta polypeptide | 1924 | -0.055 | 0.3318 | No |
| 68 | DLST | DLST Entrez, Source | dihydrolipoamide S-succinyltransferase (E2 component of 2-oxo-glutarate complex) | 2059 | -0.153 | 0.2930 | No |
| 69 | DLAT | DLAT Entrez, Source | dihydrolipoamide S-acetyltransferase (E2 component of pyruvate dehydrogenase complex) | 2060 | -0.153 | 0.2946 | No |
| 70 | MDH2 | MDH2 Entrez, Source | malate dehydrogenase 2, NAD (mitochondrial) | 2076 | -0.161 | 0.2917 | No |
| 71 | PDHA1 | PDHA1 Entrez, Source | pyruvate dehydrogenase (lipoamide) alpha 1 | 2175 | -0.231 | 0.2646 | No |
| 72 | IDH3B | IDH3B Entrez, Source | isocitrate dehydrogenase 3 (NAD+) beta | 2270 | -0.292 | 0.2393 | No |
| 73 | IDH3A | IDH3A Entrez, Source | isocitrate dehydrogenase 3 (NAD+) alpha | 2363 | -0.342 | 0.2152 | No |
| 74 | CS | CS Entrez, Source | citrate synthase | 2430 | -0.400 | 0.1994 | No |
| 75 | NNT | NNT Entrez, Source | nicotinamide nucleotide transhydrogenase | 2588 | -0.530 | 0.1576 | No |
| 76 | PDK1 | PDK1 Entrez, Source | pyruvate dehydrogenase kinase, isozyme 1 | 2596 | -0.535 | 0.1611 | No |
| 77 | SDHA | SDHA Entrez, Source | succinate dehydrogenase complex, subunit A, flavoprotein (Fp) | 2617 | -0.560 | 0.1608 | No |
| 78 | ETFA | ETFA Entrez, Source | electron-transfer-flavoprotein, alpha polypeptide (glutaric aciduria II) | 2767 | -0.692 | 0.1231 | No |
| 79 | ACO2 | ACO2 Entrez, Source | aconitase 2, mitochondrial | 2852 | -0.794 | 0.1060 | No |
| 80 | UQCRH | UQCRH Entrez, Source | ubiquinol-cytochrome c reductase hinge protein | 2908 | -0.852 | 0.0983 | No |
| 81 | IDH1 | IDH1 Entrez, Source | isocitrate dehydrogenase 1 (NADP+), soluble | 3060 | -1.087 | 0.0641 | No |
| 82 | CYCS | CYCS Entrez, Source | cytochrome c, somatic | 3167 | -1.278 | 0.0454 | No |
| 83 | ATP5I | ATP5I Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit E | 3378 | -2.426 | 0.0072 | No |
