EEB BootCamp 2020
Jump to navigation
Jump to search
| Lyme Disease (Borreliella) | CoV Genome Tracker | Coronavirus evolutuon |
|---|---|---|
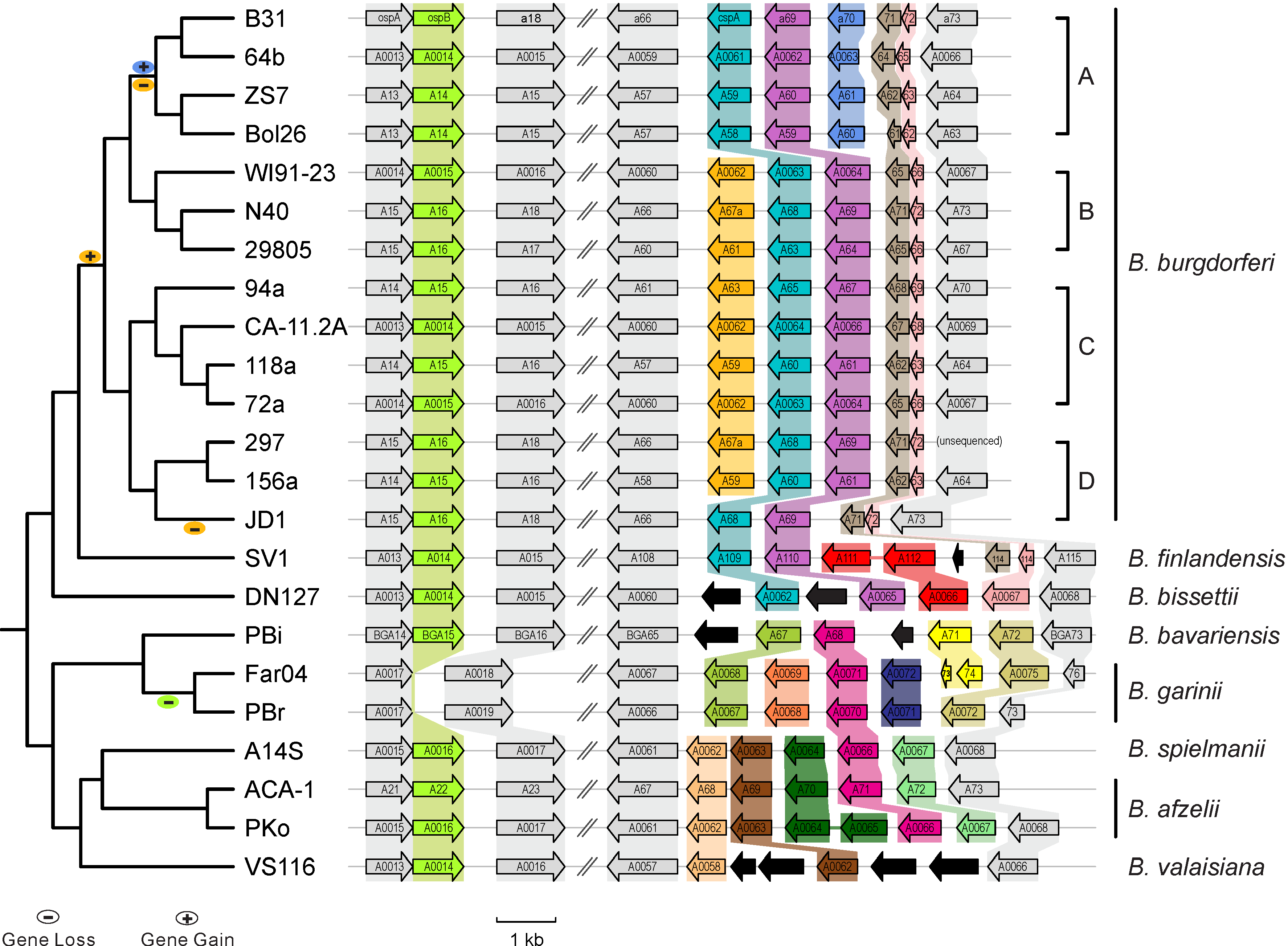
Case studies from Qiu Lab
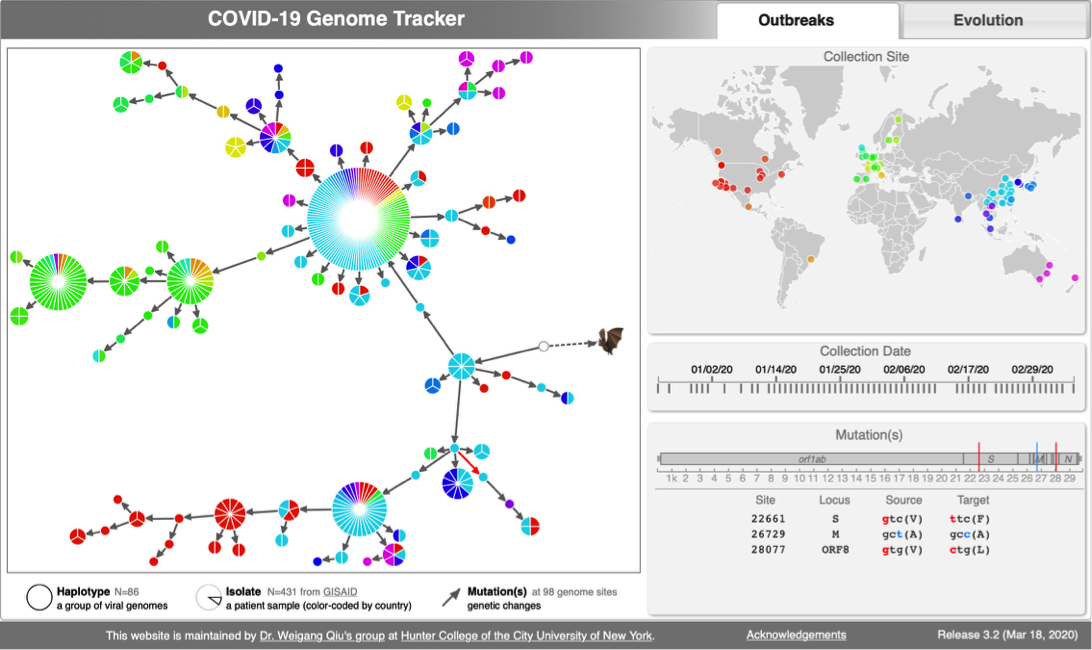
CoV genome data set
- N=565 SARS-CoV-2 genomes collected during January & February 2020. Data source & acknowledgement GIDAID (Warning: You need to acknowledge GISAID if you reuse the data in any publication)
- Download file: data file
- Create a directory, unzip, & un-tar
mkdir QiuAkther
mv cov-camp.tar.gz QiuAkther/
cd QiuAkther
tar -tzf cov-camp.tar.gz # view files
tar -xzf cov-camp.tar.gz # un-zip & un-tar
- View files
file TCS.jar
ls -lrt # long list, in reverse timeline
less Jan-Feb.mafft # an alignment of 565 CoV2 genomes in FASTA format; "q" to quit
less cov-565strains-617snvs.phy # non-gapped SNV alignment in PHYLIP format
wc hap.txt # geographic origins
head hap.txt
wc group.txt # color assignment
cat group.txt
less cov-565strains.gml # graph file (output)
Bioinformatics tools for genomic epidemiology
- BpWrapper: command-line tools for manipulation of sequences, alignment, and tree (based on BioPerl).
- Pairwise genome alignment with MUMMER: Github link
- Multiple alignment with MAFFT: Github link
- Extract SNVs with snp-sites: Github link
- Haplotype network with TCS PubMed link
- Web-interactive visualization with D3js
Tutorial
- 2-2:30: Introduction on pathogen phylogenomics
- 2:30-2:45: Demo: sequence manipulation with BpWrapper
bioseq --man
bioseq -n Jan-Feb.mafft
bioaln --man
bioaln -n -i'fasta' Jan-Feb.mafft
bioaln -l -i'fasta' Jan-Feb.mafft
bioaln -n -i'phylip' cov-565strains-617snvs.phy
bioaln -l -i'phylip' cov-565strains-617snvs.phy
FastTree -nt cov-565strains-617snvs.phy > cov.dnd
biotree --man
biotree -n cov.dnd
biotree -l cov.dnd
- 2:45-3:00: build haplotype network with TCS
# Data pre-processing
# 1. Download genomes & meta data from GISAID
# 2. Run dnadist against a reference genome
(to be added)
# 3. Remove mis-assembled and reverse-complemented genomes
bioseq -d'file:'
# 4. Remove genomes with more than 10 non-ATCG bases
bioseq -d'ambig:10'
# 5. Run mafft (not run; takes too long)
# 6. Run snp-sites
snp-sites
java -jar -Xmx1g TCS.jar
- 3:00-3:15: interactive visualization with BuTCS
- Load graph file
- Load group file
- Load haplotype file
- 3:15-3:30: Q & A